| Trees | Indices | Help |
|
|---|
|
|

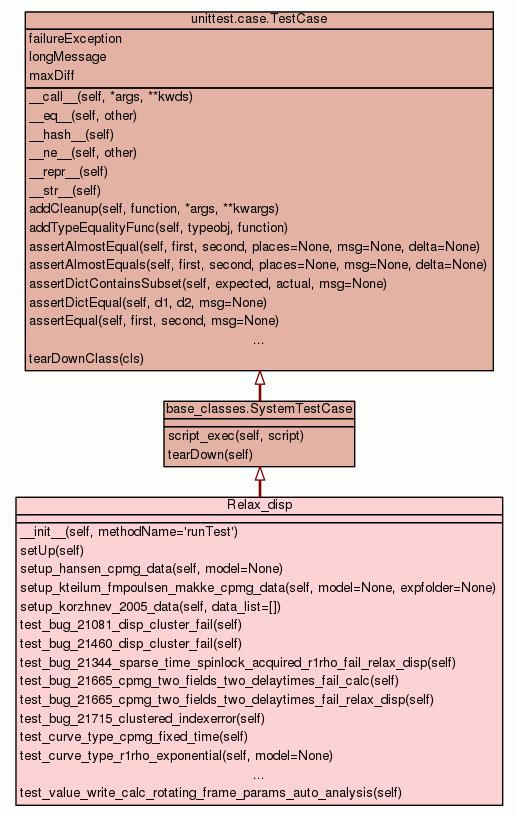
Class for testing various aspects specific to relaxation dispersion curve-fitting.
|
|||
|
Inherited from |
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
Inherited from Inherited from Inherited from Inherited from |
|||
|
|||
|
Inherited from |
|||
|
|||
|
Inherited from Inherited from |
|||
|
|||
|
Inherited from |
|||
|
|||
Skip certain tests if the C modules are non-functional.
|
Set up for all the functional tests.
|
Set up the data for the test_hansen_cpmg_data_*() system tests.
|
Set up the data for the test_kteilum_fmpoulsen_makke_cpmg_data_*() system tests.
|
Set up the data for the test_korzhnev_2005_data_*() system tests using the 'NS MMQ 2-site' model. This loads the proton-heteronuclear SQ, ZQ, DQ, and MQ (MMQ) data from:
It consists of the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant.
|
Test the curve type detection using the Dr. Flemming Hansen's CPMG fixed time test data. |
Conversion of Dr. Flemming Hansen's CPMG R2eff values into input files for CATIA. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Test of the dispersion auto-analysis using Dr. Flemming Hansen's CPMG data. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Test of the numeric model only dispersion auto-analysis using Dr. Flemming Hansen's CPMG data. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Test of the dispersion auto-analysis using Dr. Flemming Hansen's CPMG data (using the R2eff data directly instead of peak intensities). This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Test of the dispersion auto-analysis using Dr. Flemming Hansen's CPMG data with parts missing. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the CR72 dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the CR72 full dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the IT99 dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the LM63 dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the 'NS CPMG 2-site 3D' dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the 'NS CPMG 2-site 3D full' dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the 'NS CPMG 2-site expanded' dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the 'NS CPMG 2-site star' dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of Dr. Flemming Hansen's CPMG data to the 'NS CPMG 2-site star full' dispersion model. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Conversion of Dr. Flemming Hansen's CPMG R2eff values into input files for CPMGFit. This uses the data from Dr. Flemming Hansen's paper at http://dx.doi.org/10.1021/jp074793o. This is CPMG data with a fixed relaxation time period. |
Optimisation of the Korzhnev et al., 2005 15N DQ CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here only the 15N DQ data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of the Korzhnev et al., 2005 15N MQ CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here only the 15N MQ data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of the Korzhnev et al., 2005 15N SQ CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here only the 15N SQ data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of the Korzhnev et al., 2005 15N ZQ CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here only the 15N ZQ data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of the Korzhnev et al., 2005 1H MQ CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here only the 1H MQ data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of the Korzhnev et al., 2005 1H SQ CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here only the 1H SQ data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of all the Korzhnev et al., 2005 CPMG data using the 'NS MMQ 2-site' model. This uses the data from Dmitry Korzhnev's paper at DOI: 10.1021/ja054550e. This is the 1H SQ, 15N SQ, ZQ, DQ, 1H MQ and 15N MQ data for residue Asp 9 of the Fyn SH3 domain mutant. Here all data will be optimised. The values found by cpmg_fit using just this data are:
|
Optimisation of Kaare Teilum, Flemming M Poulsen, Mikael Akke 2006 "acyl-CoA binding protein" CPMG data to the CR72 dispersion model. This uses the data from paper at http://dx.doi.org/10.1073/pnas.0509100103. This is CPMG data with a fixed relaxation time period. Experiment in 0.48 M GuHCl (guanidine hydrochloride). |
Optimisation of Kaare Teilum, Flemming M Poulsen, Mikael Akke 2006 "acyl-CoA binding protein" CPMG data to the CR72 dispersion model. This uses the data from paper at http://dx.doi.org/10.1073/pnas.0509100103. This is CPMG data with a fixed relaxation time period. Experiment in 0.48 M GuHCl (guanidine hydrochloride). |
Optimisation of Kaare Teilum, Flemming M Poulsen, Mikael Akke 2006 "acyl-CoA binding protein" CPMG data to the CR72 dispersion model. This uses the data from paper at http://dx.doi.org/10.1073/pnas.0509100103. This is CPMG data with a fixed relaxation time period. Experiment in 0.48 M GuHCl (guanidine hydrochloride). Figure 3 shows the ln( k_a [s^-1]) for different concentrations of GuHCl. The precise values are:
|
Optimisation of Kaare Teilum, Flemming M Poulsen, Mikael Akke 2006 "acyl-CoA binding protein" CPMG data to the CR72 dispersion model. This uses the data from paper at http://dx.doi.org/10.1073/pnas.0509100103. This is CPMG data with a fixed relaxation time period. Experiment in 1.01 M GuHCl (guanidine hydrochloride). The comparison is to Figure 2, which is for dataset with 1 M GuHCl. The reported results are expected to be in rad.s^-1. Conversion into relax stored values is preferably. Representative 15N CPMG relaxation dispersion curve measured on the cross peaks from residue L61 in folded ACBP at pH 5.3, 1 M GuHCl, and 40C:
Conversion of paper results to relax results is performed by:
Figure 3 shows the ln( k_a [s^-1]) for different concentrations of GuHCl. The precise values are:
|
Optimisation of the Kjaergaard et al., 2013 Off-resonance R1rho relaxation dispersion experiments using the 'DPL' model. This uses the data from Kjaergaard's paper at DOI: 10.1021/bi4001062. |
Test the 'MMQ CR72' model fitting against Remco Sprangers' ClpP data. This uses the data from Remco Sprangers' paper at http://dx.doi.org/10.1073/pnas.0507370102. This is MMQ CPMG data with a fixed relaxation time period. |
Test the 'NS MMQ 2-site' model fitting against Remco Sprangers' ClpP data. This uses the data from Remco Sprangers' paper at http://dx.doi.org/10.1073/pnas.0507370102. This is MQ CPMG data with a fixed relaxation time period. |
System test of the value.write function to write intensities for an R1rho setup. This system test is to make sure, that modifying the API for special parameters theta and w_eff does not alter the functionality value.write. This uses the data of the saved state attached to bug #21344. |
System test of the value.write function to write return values of theta from calc_rotating_frame_params() function for an R1rho setup. This uses the data of the saved state attached to bug #21344. |
System test of the value.write function to write return values of w_eff from calc_rotating_frame_params() function for an R1rho setup. This uses the data of the saved state attached to bug #21344. |
System test of the auto_analysis value.write function to write theta and w_eff values for an R1rho setup. This uses the data of the saved state attached to bug #21344. |
| Trees | Indices | Help |
|
|---|
| Generated by Epydoc 3.0.1 on Mon Mar 17 15:11:55 2014 | http://epydoc.sourceforge.net |